Crop production and yield depend largely upon photosynthesis and nutrient acquisition in the face of daily and seasonal environmental fluctuations regarding water, temperature, light, and nutrient supply. In non-cultivated wild plants, e.g. wild barley (Hordeum spontaneum), investing available resources into maintaining reproductive fitness (production of viable seeds or grains) ensures population viability over time, which under limited resources is generally at the expense of yield (i.e. total seed weight per field area). The strategic challenge for plant breeders is to maximize yield (and quality) under input sustainability restraints while minimizing unpredictability and losses due to environmental fluctuations. However, environmental stresses (e.g. water availability, temperature extremes, nitrogen shortage), exacerbated by climate change, are of increasing concern for sustainable crop production.
Source: Klaus Pillen
Plant perception and responses to the external environment are mediated by molecular signaling and physiological circuits underpinned by genetic control networks that interact with plant development, architecture and phenology. In wild and cultivated barley, natural evolution and human activities, respectively, resulted in selection of beneficial pathways, genes, and alleles that have allowed barley to adapt to a wide range of environments, from the polar circle to the boundaries of deserts. While some genes and gene networks underpinning adaptive responses have been elucidated (e.g. flowering time), most remain poorly understood. Thanks to the recently achieved availability of high-quality full genome sequences and new genetic tools, stress response networks can now be clarified directly in crop species like barley, paving the link between stress response and agricultural yield.
Source: Klaus Pillen
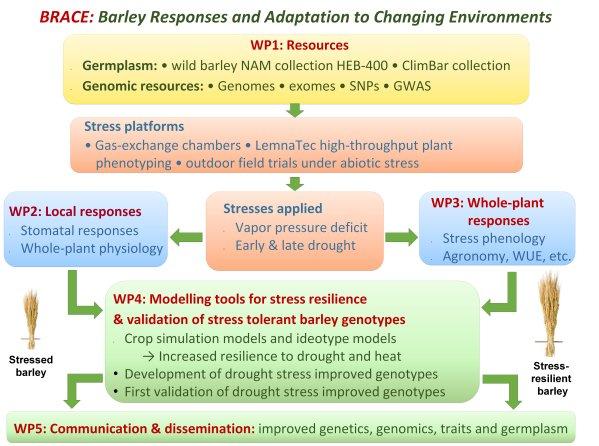
The BRACE consortium aims to resolve the components of abiotic stress responses and dynamics and to identify genes and alleles needed for resilience, by using specialized populations of wild and cultivated barley as well as genotypes containing mutated candidate genes for stress resilience. The highly divergent wild x cultivated barley nested association mapping (NAM) population HEB-25 contains genes and alleles needed for abiotic stress resilience, as shown in a total of 19 so far published reports. In BRACE we plan to test if particular wild barley alleles can be identified and selected to improve resilience to drought under test conditions as well as under field conditions and if these alleles can be successfully introgressed into locally adapted elite barley cultivars. A manageable set of 400 wild barley HEB lines (termed HEB-400) has been selected and will be deeply characterized on advanced environmental and phenotyping platforms that include gas-exchange chambers, high-throughput greenhouses, and multi-location experimental field trials.
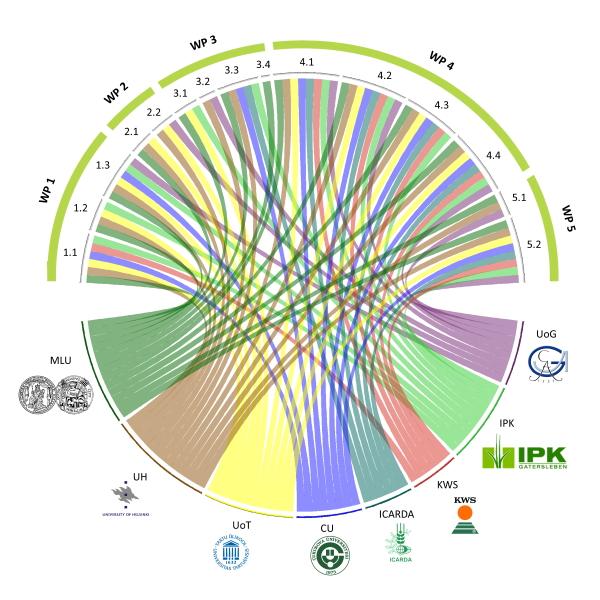
In a multi-disciplinary approach, responses to drought stress will be analyzed biochemically, physiologically, genetically, genomically, and through crop simulation modelling, with reference to gene function and allele diversity. Information on pathways, allele diversity, and crop simulation and ideotype models will be integrated to reveal mechanisms and deliver genes and alleles from wild barley growth models and improved prediction models of barley performance cultivated under abiotic stresses, such as drought. Public and private breeding organizations, included in BRACE and located in European and in wider barley cultivation areas (i.e. Turkey and North Africa), will co-exploit improved germplasm and the derived knowledge to develop cultivars better adapted to future climate conditions.

Source: Klaus Pillen
Prof. Klaus Pillen
Martin-Luther-University Halle-Wittenberg, GERMANY
Email: klaus.pillen@landw.uni-halle.de
Prof. Alan Schulman
University of Helsinki, FINLAND
Prof. Hannes Kollist
University of Tartu, ESTONIA
Prof. Hakan Ozkan
Cukurova University, Faculty of Agriculture, TURKEY
Dr. Michael Baum
International Center for Agricultural Research in the Dry Areas (ICARDA), LEBANON
Dr. Klaus Oldach
KWS LOCHOW GMBH, GERMANY
Dr. Kerstin Neumann
Leibniz Institute of Plant Genetics and Crop Plant Research (IPK), GERMANY
Prof. Reimund Rötter
University of Göttingen, GERMANY
Dr. Paul Shaw (Associate partner)
James Hutton Institute, UK
Dr. Luigi Cattivelli
Research Centre for Genomics and Bioinformatics (CREA), ITALY
Dr. Max Schulman
Central Union of Agricultural Producers and Forest Owners (MTK), Finland
Dr. Ahmad M. Alqudah
Aarhus University, Denmark